This represented a major change in the Cytoscape architecture it is a more modularized, expandable and maintainable version of the software. The Cytoscape core developer team continues to work on this project and released Cytoscape 3.0 in 2013. In this session, we will learn about new capabilities to integrate Cytoscape into programmatic workflows and pipelines using R. Version 3.0 was released Feb 1, 2013, and the latest version, 3.4.0, was released in May 2016. Cytoscape (is probably the most popular applications for network analysis and visualization with GUI. Workshop Materials PowerPoints Part-1 GEO Data Mining () Part-2 Pathway Enrichment Analysis - g:Profiler and GSEA () Part-3 Cytoscape - Data Visualization () Hands-On Exercise - differential gene expression dataset for SARS-Cov2 infected vs. Version 2.0 was initially released in 2004 Cytoscape 2.83, the final 2.xx version, was released in May 2012. Version 1.1.1 is the last stable release for the 1.0 series. 
Cytoscape was initially made public in July, 2002 (v0.8) the second release (v0.9) was in November, 2002, and v1.0 was released in March 2003. Prepare data Go to the RCSB PDB Protein Comparison Tool web site ( Enter 4hhb in the text field for PDB1 and the tool will automatically select chain A (. Now, it is developed by an international consortium of open source developers. Cytoscape also has a JavaScript-centric sister project named Cytoscape.js that can be used to analyse and visualise graphs in JavaScript environments, like a browser.Ĭytoscape was originally created at the Institute of Systems Biology in Seattle in 2002. Plugins may be developed using the Cytoscape open Java software architecture by anyone and plugin community development is encouraged. Plugins are available for network and molecular profiling analyses, new layouts, additional file format support and connection with databases and searching in large networks. Additional features are available as plugins. Just put a URL to it here and well apply it, in the order you have them, before the CSS in. Several case studies of Cytoscape plug-ins are surveyed, including a search for interaction pathways correlating with changes in gene expression, a study of protein complexes involved in cellular recovery to DNA damage, inference of a combined physical/functional interaction network for Halobacterium, and an interface to detailed stochastic/kinetic gene regulatory models.Cytoscape is an open source bioinformatics software platform for visualizing molecular interaction networks and integrating with gene expression profiles and other state data. You can apply CSS to your Pen from any stylesheet on the web. 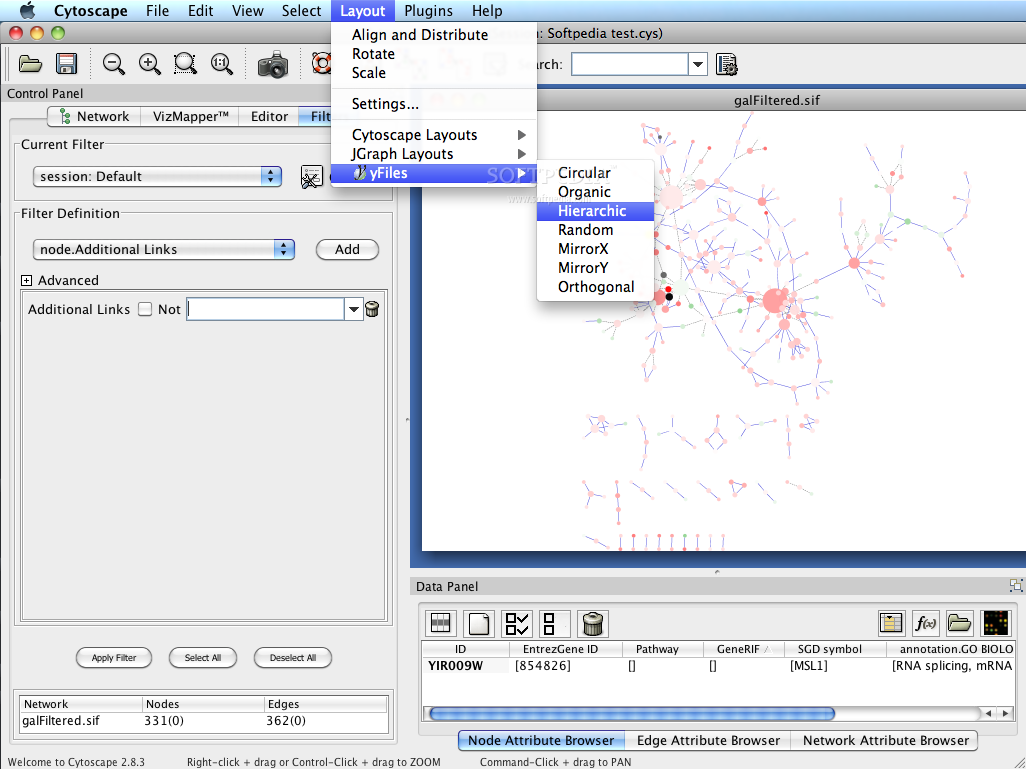
The Core is extensible through a straightforward plug-in architecture, allowing rapid development of additional computational analyses and features. Cytoscape's software Core provides basic functionality to layout and query the network to visually integrate the network with expression profiles, phenotypes, and other molecular states and to link the network to databases of functional annotations. Although applicable to any system of molecular components and interactions, Cytoscape is most powerful when used in conjunction with large databases of protein-protein, protein-DNA, and genetic interactions that are increasingly available for humans and model organisms. Cytoscape is an open source software project for integrating biomolecular interaction networks with high-throughput expression data and other molecular states into a unified conceptual framework.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
